The Spillover Spreading Matrix can be displayed for any type of compensation matrix by selecting the “SSM” button above the compensation matrix in the compensation window.ĪutoSpill: a method for calculating spillover coefficients in high-parameter flow cytometry.Īuthors: Carlos P. When calculating the Spillover Spreading Matrix for AutoSpread matrices, FlowJo will automatically use the AutoSpread algorithm, which applies a linear model to derive the values. 
The traditional Spillover Spreading Calculation requires a defined positive and negative population to calculate the Spillover Spreading values. The AutoSpill compensation method requires a modified algorithm for calculating the Spillover Spreading Matrix (SSM). The spillover signal from this unstained control into the other detectors will then be accounted for during the AutoSpill calculation allowing the algorithm to remove the effects of autofluorescence in the final matrix. An unstained control can be added to an empty detector (a detector with no stain associated with it) when setting up the compensation matrix in the compensation wizard. One advantage of the AutoSpill compensation method is that it is able to remove the autofluorescence associated with the cells using an unstained control as a baseline. The resulting AutoSpill compensation will appear the same as any traditional compensation matrix with a grid of values indicating the compensation being applied. The autospill algorithm will use that population’s parent, which is the “Size” gate. For example, in the image below the population “APC-H7-A+” is listed in the positive column. Instead, the autospill algorithm will take the parent population of the population displayed in the “Positive” column and it will use that parent population to generate results. Note: The “Positive” column will still display a positive population, even though Autospill will not use this population. Once the compensation is properly set up the “View Matrix” button can be selected to generate the AutoSpill compensation matrix. Additionally the AutoSpill compensation does not require universal negative controls so the “Remove Univ Neg” button was created to quickly remove the negative control that FlowJo will automatically add. A size gate that removes outliers is still essential for isolating representative events, and the size gate drawn by FlowJo may need adjustment to contain the representative populations. Unlike traditional compensation, the AutoSpill method does not require gated positive and negative populations, as the algorithm will determine its own positive populations. 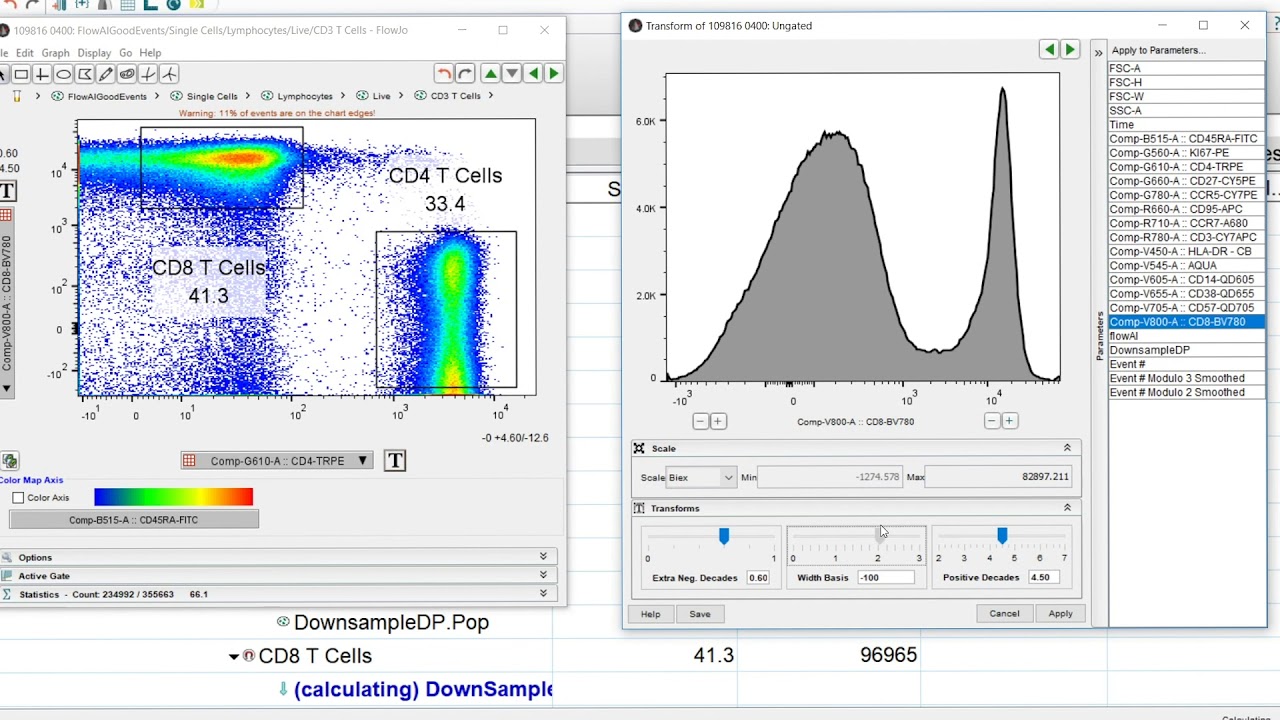
FlowJo may also try to load an unstained control for use as a universal negative. Compensation controls must first be loaded into the FlowJo compensation group, then the compensation button at the top of the workspace can be selected.įlowJo will automatically try to match the compensation controls to the appropriate detectors, apply a size gate to each of the controls, then try to gate the positive and negative populations in each control. Performing AutoSpill compensation has similar setup to regular compensation. The option to return to traditional compensation is located next to the AutoSpill tick box and there is also a button for removing the universal negative, which is not required for the AutoSpill algorithm but can be used to remove autofluorescence. When the option is ticked FlowJo will use the AutoSpill algorithm to calculate the compensation matrix. The AutoSpill option can be found in the compensation wizard. Additionally the AutoSpill function can be used to remove the signal due to autofluorescence. The new algorithm, combines automatic gating with linear regression through an iterative process that helps reduce error and improve the compensation matrix. This form of gate-less compensation removes some of the requirements of traditional compensation making compensation faster and simpler. AutoSpill is a new algorithm for calculating spillover and producing a compensation matrix developed and incorporated into FlowJo v10.7 in collaboration with researchers from the Vlaams Instituut voor Biotechnologie in Belgium and the the Babraham Institute in Cambridge, UK.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
